Output directory structure
Author: Menachem Sklarz
Table of Contents
The main directory structure
The directories are elaborated on below.
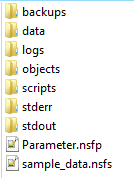
The directory structure created by NeatSeq-Flow
The scripts directory
Executing
bash 00.workflow.commands.shwill execute the entire workflow.The scripts beginning
01.Import…etc. execute entire steps.The actual scripts running each step per sample or on the entire project are contained in the equivalent directories
01.Import…etc.The scripts are numbered by execution order (see
00.workflow.commands.sh)
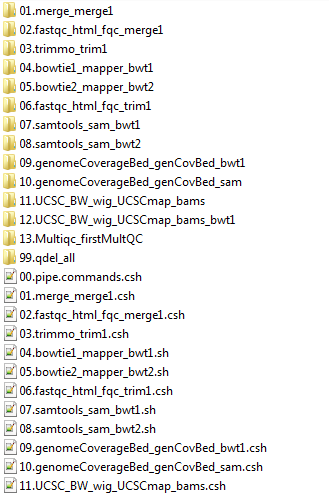
The scripts directory created by NeatSeq-Flow
The data directory
In the data directory, the analysis outputs are organized by module, by module instance and by sample.
Below is the data directory for the example, showing the tree organization for the bowtie2_mapper and Multiqc modules.
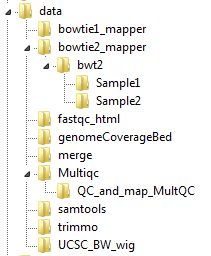
The data directory created by NeatSeq-Flow, showing the tree organization.
The backup directory
The backup directory contains a history of workflow sample and parameter files.
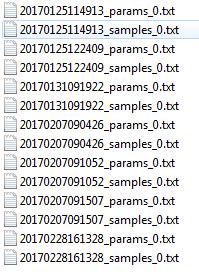
The backups directory created by NeatSeq-Flow
The logs directory
The logs directory contains various logging files:
version_list. A list of all the versions of the workflow with equivalent comments
file_registration. A list of files produced, including md5 signatures, and the script and workflow version that produced them
log_file_plotter.R. An R script for producing a plot of the execution times. (Run with Rscript and receives a single argument – a log file to plot)log_<workflow_ID>.txt. Log of the execution times of the script per workflow version ID.log_<workflow_ID>.txt.html. Graphical representation of the progress of the WF execution, as produced by thelog_file_plotter.Rscript (see figure below)
The logs directory created by NeatSeq-Flow
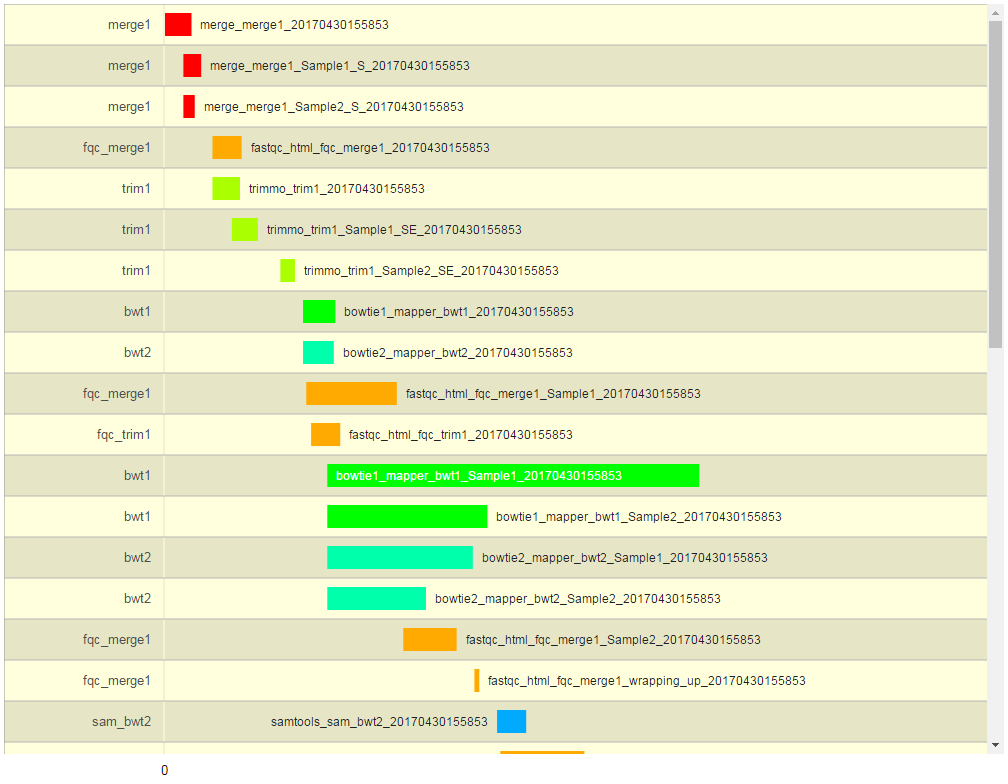
A graphical representation of the workflow execution.
The stderr and stdout directories
The stderr and stdout directories store the script standard error and outputs, respectively.
These are stored in files containing the module name, module instance, sample name, workflow ID and cluster job ID.
The objects directory
The objects directory contains various files describing the workflow:
The objects directory created by NeatSeq-Flow
pipeline_graph.html: An SVG diagram of the workflow.diagrammer.R: an R script for producing a DiagrammeR diagram of the workflow.pipedata.json: A JSON file containing all the workflow data, for uploading to JSON compliant databases etc.workflow_graph.htmlis the output from executingRscript diagrammer.R.