NeatSeq-Flow: A Lightweight Software for Efficient Execution of High-Throughput Sequencing Workflows.¶


Important links
- Quick Start: install and try NeatSeq-Flow
- GitHub: NeatSeq-Flow
- GitHub: NeatSeq-Flow GUI
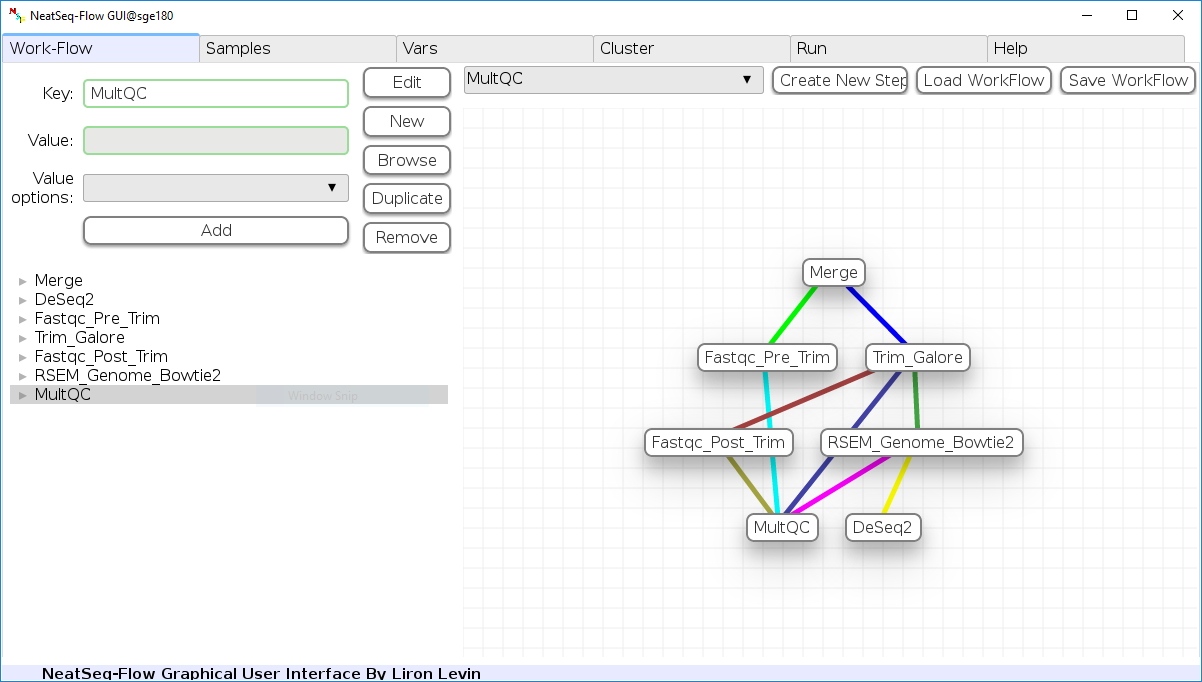
- A short movie demonstrating the GUI
- Module and Workflow repository
- Article on BioRXiv
- Contact
What is NeatSeq-Flow?¶
NeatSeq-Flow is a platform for modular design and execution of bioinformatics workflows on a local computer or, preferably, computer cluster. The platform has a command-line interface as well as a fully functional graphical user interface (GUI), both used locally without the need to connect to remote servers. Analysis programs comprising a workflow can be anything executable from the Linux command-line, either publicly available or in-house programs. Ready-to-use workflows are available for common Bioinformatics analyses such as assembly & annotation, RNA-Seq, ChIP-Seq, variant calling, metagenomics and genomic epidemiology. Creation and sharing of new workflows is easy and intuitive, without need for programming knowledge. NeatSeq-Flow is general-purpose and may easily be adjusted to work on different types of analyses other than high-throughput sequencing.
NeatSeq-Flow is fully accessible to non-programmers, without compromising power, flexibility and efficiency. The user only has to specify the location of input files and the workflow design, and need not bother with the location of intermediate and final files, nor with transferring files between workflow steps. Workflow execution is fully parallelized on the cluster, and progress can be inspected through NeatSeq-Flow “terminal monitor”. All workflow steps, parameters and order of execution are stored in one file, which together with the shell scripts produced by NeatSeq-Flow comprise a complete documentation of the workflow and enable future execution of the exact same workflow or modifications thereof.
Available Modules and Workflows¶
Quick Start:¶
- Installing Using Conda will install NeatSeq-Flow with all its dependencies in one go:
First if you don’t have Conda, install it!
Then in the terminal:
- Create the NeatSeq_Flow conda environment:
conda env create levinl/neatseq_flow
- Activate the NeatSeq_Flow conda environment:
bash source activate NeatSeq_Flow
- Run NeatSeq_Flow_GUI:
NeatSeq_Flow_GUI.py --Server
Use the information in the terminal:
- Managing Users:
- It is possible to mange users using SSH, NeatSeq-Flow will try to login by ssh to a host using the provided “User Name” and “Password”.
- The ssh host can be local or remote.
- Note: If using a remote host, NeatSeq-Flow needs to be installed on the remote host and the analysis will be run on the remote host by the user that logged-in
NeatSeq_Flow_GUI.py --Server --SSH_HOST 127.0.0.1
- For more option:
NeatSeq_Flow_GUI.py -h
Authors¶
- Menachem Sklarz
- Liron Levin
- Michal Gordon
- Vered Chalifa-Caspi
Bioinformatics Core Facility, llse Katz Institute for Nanoscale Science and Technology, Ben-Gurion University of the Negev, Beer-Sheva, Israel
Cite NeatSeq-Flow¶
Contact Us¶
Web Site Contents:¶
First Steps